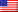
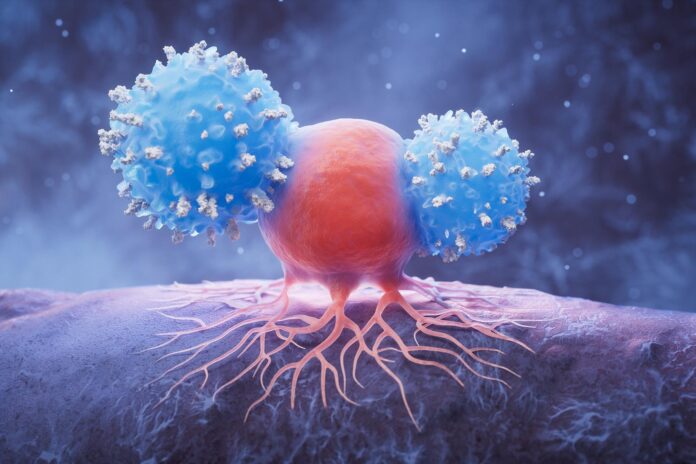
Cancer is often described as a genetic lottery, where different patients develop tumors driven by unique, complex mutations. Traditionally, this has meant that treating cancer requires a highly personalized, mutation-by-mutation approach—a strategy that is scientifically impressive but clinically difficult to scale.
However, a new study published in Nature suggests a paradigm shift. Researchers have developed a technology called PerturbFate that reveals how hundreds of different genetic errors often converge on the same few cellular “control switches.” By identifying these shared downstream targets, scientists may be able to design simpler, more universal therapies that work across a wide variety of genetic causes.
The Problem of Genetic Complexity
Modern genomics has successfully identified thousands of mutations linked to diseases like cancer and neurodegenerative disorders. Yet, this knowledge has not translated into equally effective treatments.
The core challenge is diversity. Mutations often occur in different biological pathways—some affect how genes are read, others alter cell signaling or structure. Because these errors look so different, they are hard to target with a single drug. As a result, therapies often fail when they only address one specific mutation while ignoring the broader cellular chaos.
Junyue Cao, head of the Laboratory of Single-Cell Genomics and Population Dynamics, posed a critical question: Do these mutations truly act independently?
“We wondered whether all these different genes may be mediated by some shared downstream signaling that we can discover and target instead,” says Cao.
If many distinct mutations funnel into the same final cellular behavior, then treating that common behavior—not the individual mutations—could be a more effective strategy.
Introducing PerturbFate
To test this hypothesis, the team needed a tool capable of tracking how genetic changes reshape cells in real time. Existing methods often provided only a snapshot of one molecular layer (like gene expression) or failed to capture the dynamic changes as they happened.
Zihan Xu, a graduate student in Cao’s lab, developed PerturbFate, a single-cell platform that bridges this gap. The technology allows researchers to:
- Perturb hundreds to thousands of genes in parallel.
- Measure multiple molecular layers simultaneously in the same single cell, including DNA accessibility (chromatin state) and RNA production (gene expression).
- Track these changes over time to map how early genetic disruptions lead to later disease states.
By combining DNA and RNA data from individual cells, PerturbFate can reconstruct the entire gene regulatory network, showing exactly how different mutations lead to similar outcomes.
Case Study: Melanoma Drug Resistance
The team tested PerturbFate using melanoma drug resistance as a model. Melanoma tumors often become resistant to drugs like Vemurafenib through many different genetic routes.
The researchers selected 143 genes known to be linked to this resistance and systematically turned them off in melanoma cells. They then used PerturbFate to monitor the cellular response, analyzing over 300,000 cells.
The results were striking:
1. Convergent Pathways: Despite starting with different genetic disruptions, many of these changes pushed the cells into the same drug-resistant state.
2. Shared Regulatory Nodes: The platform identified common “control points” or regulatory nodes that drove this resistance.
3. Effective Intervention: When the researchers targeted these shared nodes, drug resistance decreased significantly.
The Mediator Complex Connection
The study also uncovered a specific biological mechanism involving the Mediator Complex, a protein system that helps control gene activity. Disrupting different parts of this complex triggered drug resistance through separate initial mechanisms. However, these paths eventually converged on the same survival signal: VEGFC.
When the team blocked the VEGFC signal, the resistant cells stopped growing. This finding demonstrates that even if the starting mutations are different, blocking the final common pathway can stop the disease progression.
Implications for Future Therapies
This research suggests that complex genetic variation does not always require equally complex treatments. Instead of trying to target every individual mutation, clinicians could focus on the shared regulatory nodes that drive disease.
This approach offers several advantages:
– Simplicity: Fewer targets mean potentially simpler drug designs.
– Broad Applicability: A single therapy could work for patients with different genetic profiles.
– Predictive Power: Understanding these networks helps predict how tumors might evolve resistance.
The team has made the PerturbFate tools publicly available and plans to expand the research beyond cultured cells into living systems. Future applications may include studying aging and Alzheimer’s disease, where similar complex genetic interactions are at play.
“This is just a starting point,” says Cao. “Now that we’ve demonstrated the approach in a simple model, we’re working to extend it into living systems to study even more complex diseases.”
Conclusion
The development of PerturbFate marks a significant step toward simplifying cancer treatment. By revealing that diverse genetic mutations often share common cellular endpoints, this technology opens the door to more unified, effective therapies that target the root of disease behavior rather than its myriad genetic causes.